Exploring biomolecular structure, stability and interactions with native MS
Photoreceptors are a widespread class of chromophore-containing proteins that mediate various biological processes triggered by the absorption of light at species-specific wavelengths.
Upon photoexcitation, intramolecular proton or electron transfer typically induces conformational changes that enable the recruitment of downstream signaling partners.
Ion Mobility Spectrometry (IMS) coupled with nano Electrospray Ionisation Mass Spectrometry (nESI-MS) is a powerful approach to investigate conformational protein dynamics. Our work focuses on the time-resolved analysis of structural rearrangements in photoactive proteins upon irradiation.
For this purpose, we implemented an irradiation setup that enables light activation directly during ionization. Light-induced transitions into different signaling states can be detected as characteristic changes in the mass spectra. In addition, IMS allows investigation of conformational transitions on millisecond-to-second timescales and enables the determination of rate constants. Experimental collision cross sections (CCS) obtained for different conformations can be correlated with CCS values calculated from structural models such as NMR or crystal structures. Furthermore, collision-induced unfolding experiments provide insights into protein stability and its relationship to conformational properties.
Photoreceptors are a widespread class of chromophore-containing proteins that mediate various biological processes triggered by the absorption of light at species-specific wavelengths.
Upon photoexcitation, intramolecular proton or electron transfer typically induces conformational changes that enable the recruitment of downstream signaling partners.
Ion Mobility Spectrometry (IMS) coupled with nano Electrospray Ionisation Mass Spectrometry (nESI-MS) is a powerful approach to investigate conformational protein dynamics. Our work focuses on the time-resolved analysis of structural rearrangements in photoactive proteins upon irradiation.
For this purpose, we implemented an irradiation setup that enables light activation directly during ionization. Light-induced transitions into different signaling states can be detected as characteristic changes in the mass spectra. In addition, IMS allows investigation of conformational transitions on millisecond-to-second timescales and enables the determination of rate constants. Experimental collision cross sections (CCS) obtained for different conformations can be correlated with CCS values calculated from structural models such as NMR or crystal structures. Furthermore, collision-induced unfolding experiments provide insights into protein stability and its relationship to conformational properties.
Heat shock proteins are important chaperones that have distinctive protective properties, such as preventing unwanted aggregation of proteins or stabilizing and even (re-)folding its client molecules. These proteins are abundant in almost all known organisms and play a key role in stress response, as they are expressed when a cell is under strain. Large parts of the human proteome are known to be client molecules of one or another Heat shock protein. Our main target of interest is the HSP90. Together with some co-chaperones it has an important role in the lifecycle of glucocorticoid receptors and protein kinases. We aim to describe the interactions between HSP90, its co-chaperons and its client molecules via native mass spectrometry. With regard to binding affinities and conformational changes, we try to clarify parts of their vital role in the lifespan of cells.
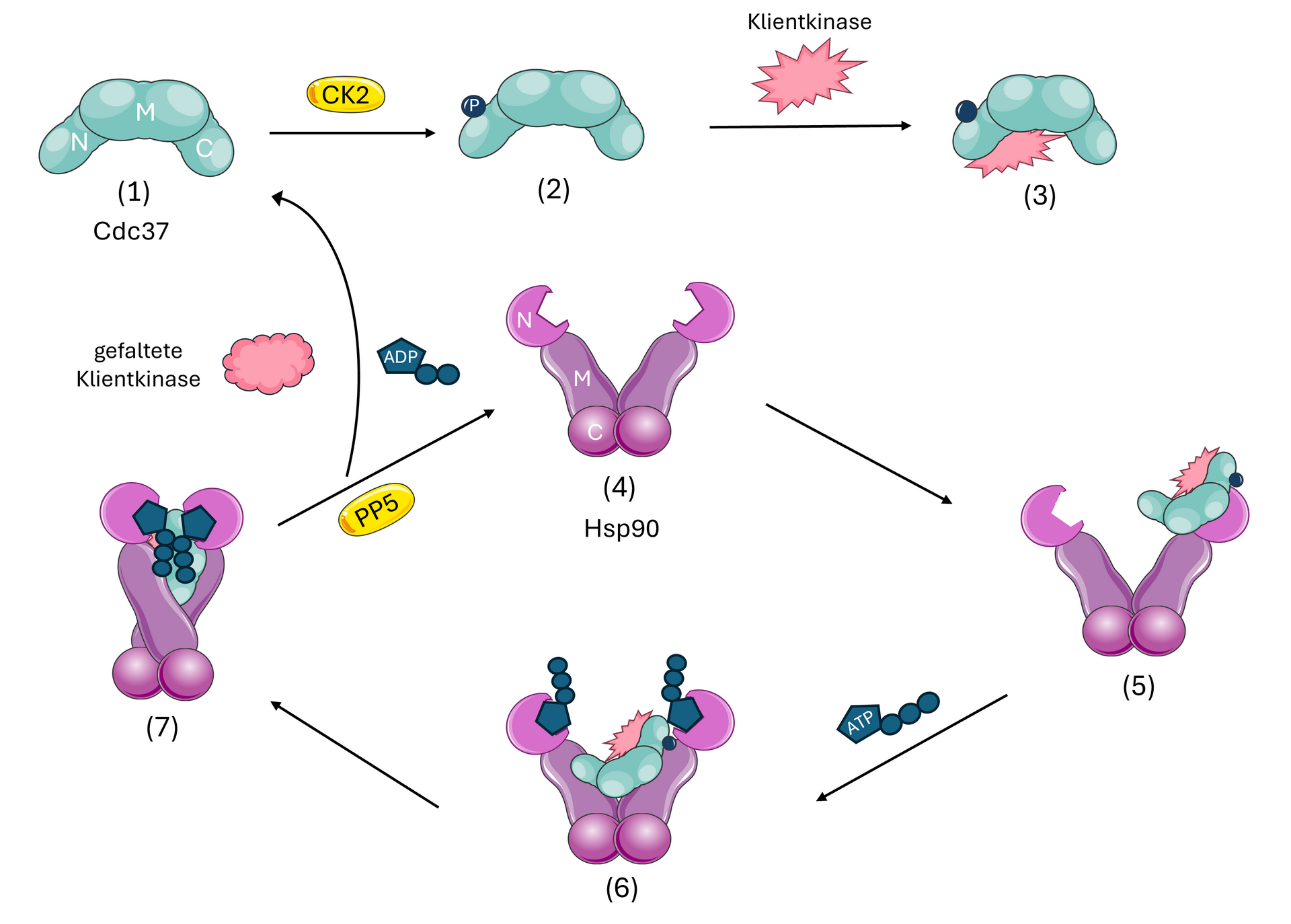
Heat shock proteins are important chaperones that have distinctive protective properties, such as preventing unwanted aggregation of proteins or stabilizing and even (re-)folding its client molecules. These proteins are abundant in almost all known organisms and play a key role in stress response, as they are expressed when a cell is under strain. Large parts of the human proteome are known to be client molecules of one or another Heat shock protein. Our main target of interest is the HSP90. Together with some co-chaperones it has an important role in the lifecycle of glucocorticoid receptors and protein kinases. We aim to describe the interactions between HSP90, its co-chaperons and its client molecules via native mass spectrometry. With regard to binding affinities and conformational changes, we try to clarify parts of their vital role in the lifespan of cells.
The
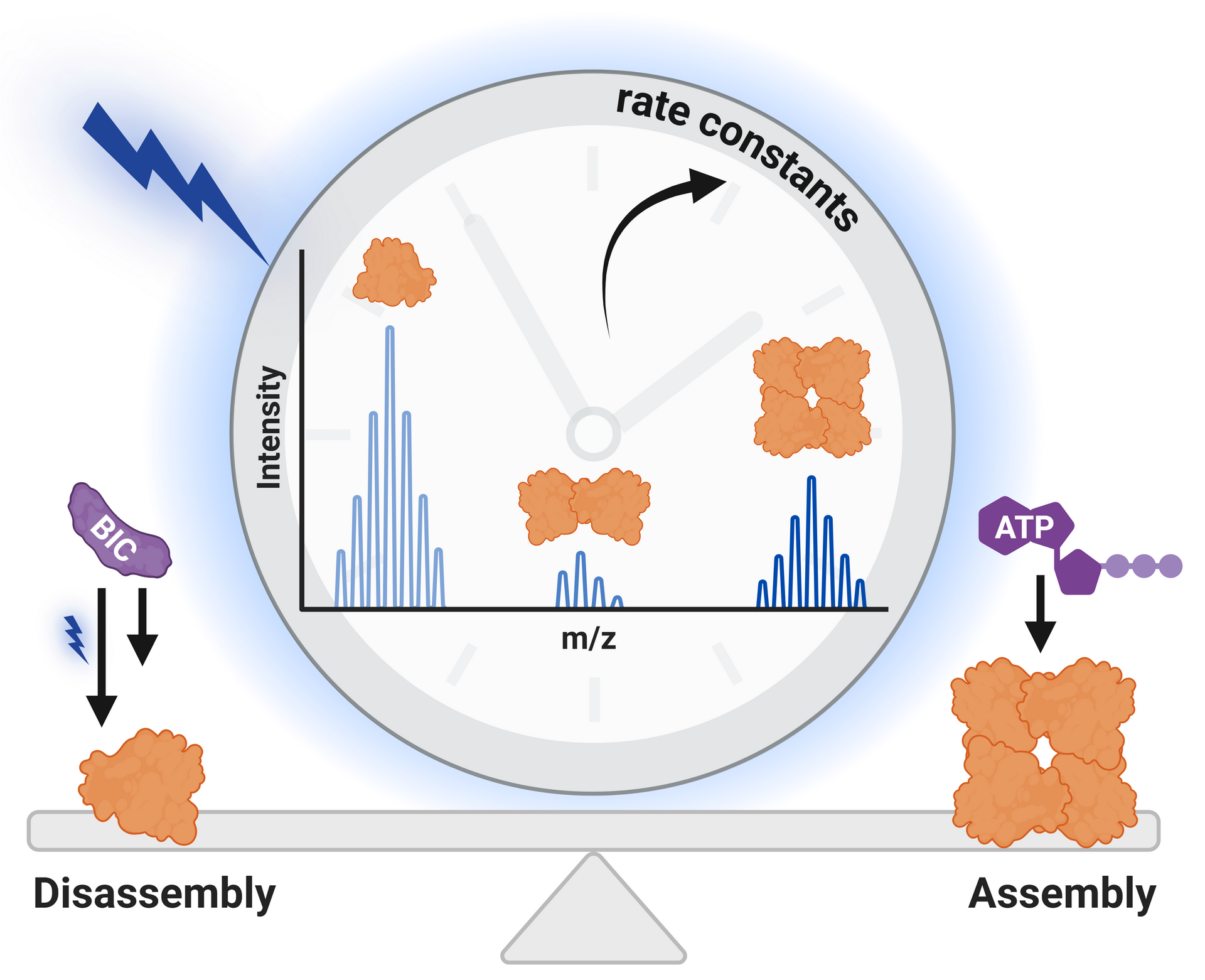
association and dissociation of non-covalently interacting species play
a vital role in various biological processes, ranging from the
formation of DNA double helices to the functional interplay of subunits
in molecular machines. To study the pathways of these reactions, it is
crucial to not only determine the interacting partners and their
stoichiometry, but to further characterize their interactions in terms
of binding affinities.
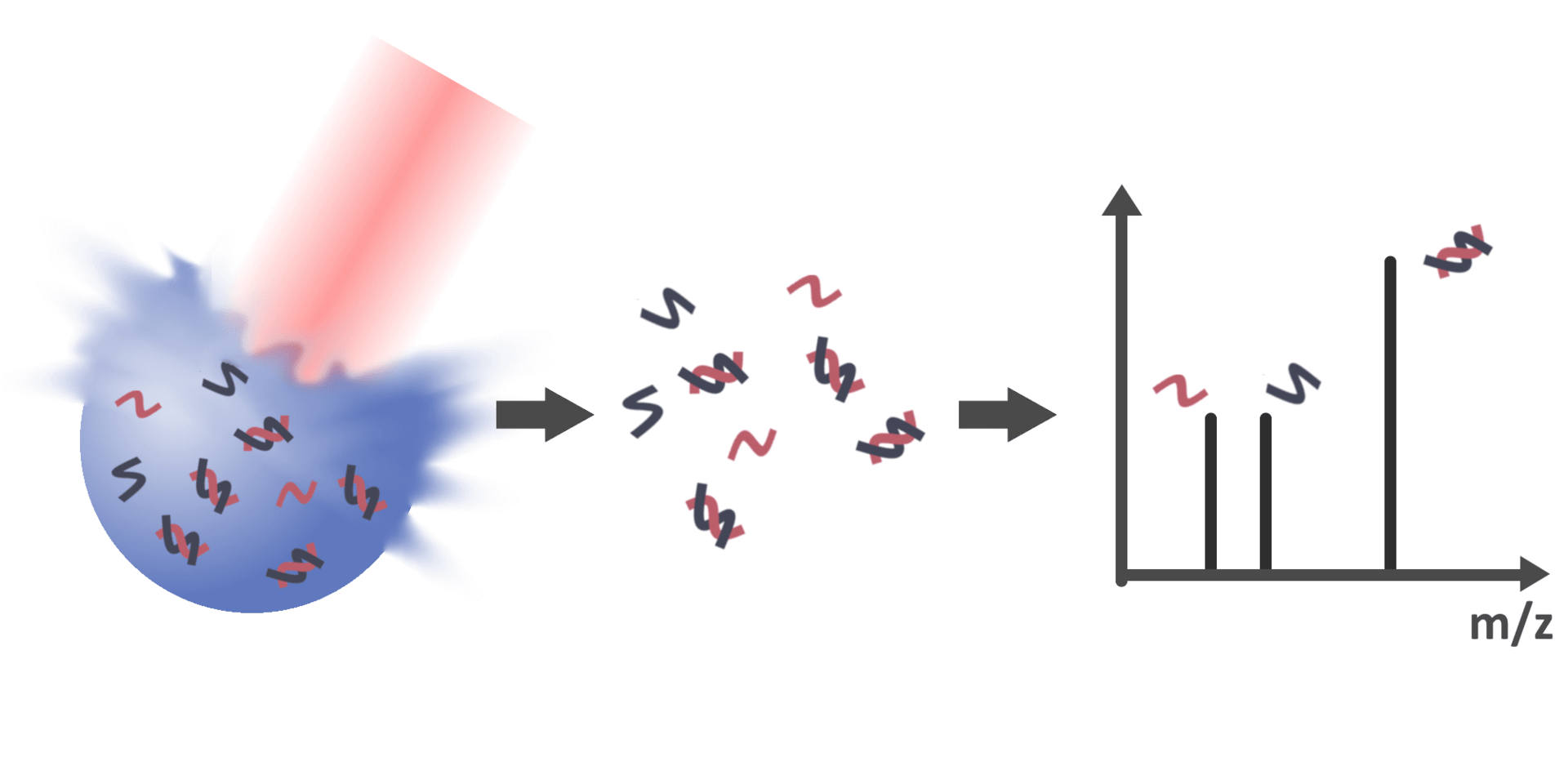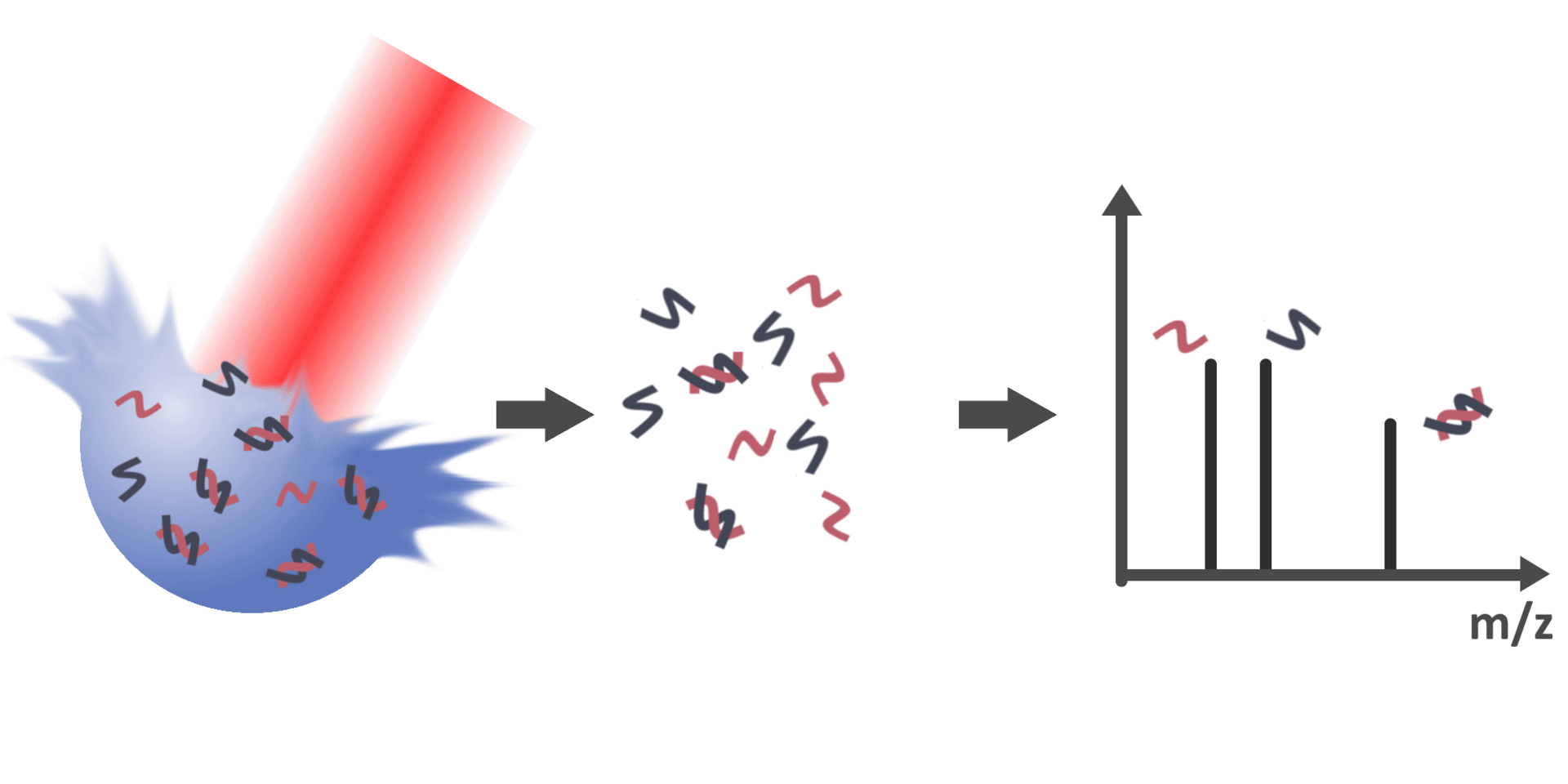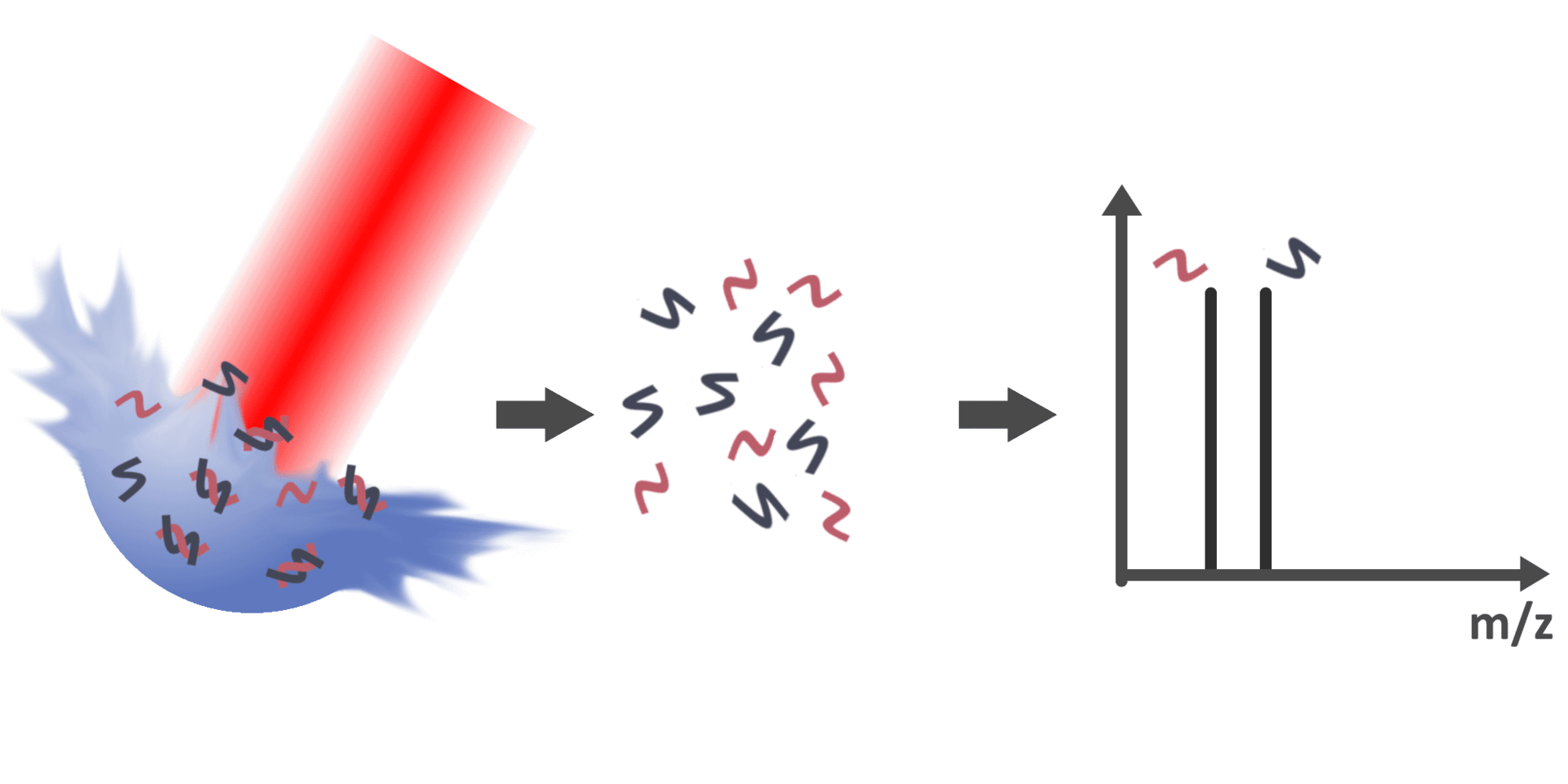
With the possibility to detect biomolecules and their non-covalently
bound complexes via LILBID-MS, the subsequent objective is therefore to
gain information about the molar quantities of bound and unbound
species in solution. However, peak areas in the mass spectra are not
directly correlated to the concentration of respective species in their
native environment due to various factors, such as dissociation of
bonds during desorption in the ion source or varying ionization
efficiencies of different species. Therefore, this project aims to
identify and account for these factors to enable the determination of
quantitative values such as the dissociation constant of specific
interactions.
The
association and dissociation of non-covalently interacting species play
a vital role in various biological processes, ranging from the
formation of DNA double helices to the functional interplay of subunits
in molecular machines. To study the pathways of these reactions, it is
crucial to not only determine the interacting partners and their
stoichiometry, but to further characterize their interactions in terms
of binding affinities.
With the possibility to detect biomolecules and their non-covalently
bound complexes via LILBID-MS, the subsequent objective is therefore to
gain information about the molar quantities of bound and unbound
species in solution. However, peak areas in the mass spectra are not
directly correlated to the concentration of respective species in their
native environment due to various factors, such as dissociation of
bonds during desorption in the ion source or varying ionization
efficiencies of different species. Therefore, this project aims to
identify and account for these factors to enable the determination of
quantitative values such as the dissociation constant of specific
interactions.
The cell interior of Gram-negative bacteria is protected by a double layer of membranes. While the inner one is mainly composed of phospholipids (PLs), the outer membrane is built as an asymmetric bilayer, where the inner part consists mainly of PLs, while the outer membrane contains lipopolysaccharides (LPS). This asymmetry plays an essential part in drug resistance, as the membrane system works as a barrier against all kinds of pharmaceutics (e.g. antibiotics). To maintain the composition of these membranes, there are PL- and LPS-transport systems in bacteria. The molecular principles that control these transport systems are the subject of our investigations - currently we are working on the Lpt- and the Mla system, which have come into focus as potential antibiotic targets. Through native mass spectrometry, in particular LILBID MS, we aim to analyse the interactions between the components of the transport systems. Furthermore, we try to follow the path a lipid or a LPS takes through the systems by binding studies.
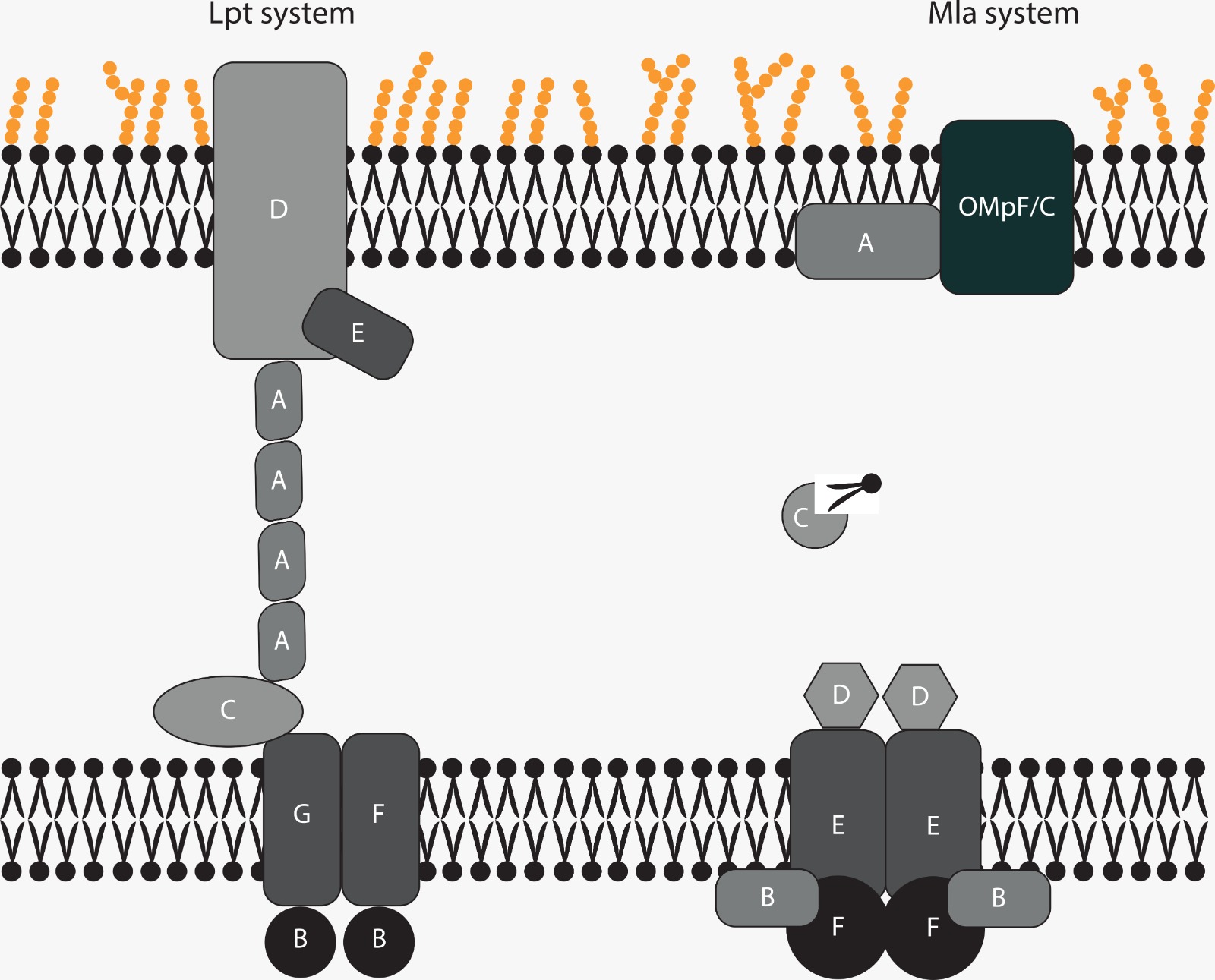
The cell interior of Gram-negative bacteria is protected by a double layer of membranes. While the inner one is mainly composed of phospholipids (PLs), the outer membrane is built as an asymmetric bilayer, where the inner part consists mainly of PLs, while the outer membrane contains lipopolysaccharides (LPS). This asymmetry plays an essential part in drug resistance, as the membrane system works as a barrier against all kinds of pharmaceutics (e.g. antibiotics). To maintain the composition of these membranes, there are PL- and LPS-transport systems in bacteria. The molecular principles that control these transport systems are the subject of our investigations - currently we are working on the Lpt- and the Mla system, which have come into focus as potential antibiotic targets. Through native mass spectrometry, in particular LILBID MS, we aim to analyse the interactions between the components of the transport systems. Furthermore, we try to follow the path a lipid or a LPS takes through the systems by binding studies.
In another ongoing project, we are integrating a LILBID ion source into a commercially available mass spectrometer platform (QTOF Ultima, Waters). The goal of this approach is to combine the unique advantages of buffer tolerance and soft ionization that LILBID-MS offers with the enhanced mass resolution, sensitivity, and extended analytical capabilities of a high-performance commercial instrument. The integration enables the use of additional instrument features such as a collision cell for controlled activation experiments, thereby expanding the accessible structural information. This hybrid setup represents an important step toward improving both data quality and methodological flexibility in native mass spectrometry applications.
In another ongoing project, we are integrating a LILBID ion source into a commercially available mass spectrometer platform (QTOF Ultima, Waters). The goal of this approach is to combine the unique advantages of buffer tolerance and soft ionization that LILBID-MS offers with the enhanced mass resolution, sensitivity, and extended analytical capabilities of a high-performance commercial instrument. The integration enables the use of additional instrument features such as a collision cell for controlled activation experiments, thereby expanding the accessible structural information. This hybrid setup represents an important step toward improving both data quality and methodological flexibility in native mass spectrometry applications.
Our group continuously develops new approaches to further improve existing methods and instrumentation for native mass spectrometry. For example, we are implementing Paul trap–based strategies to levitate microdroplets, enabling the investigation of fast, time-resolved reactions prior to ionization in LILBID-MS experiments.
In addition, we are advancing spectral quality in ESI-MS through MS-SIEVE, a setup that allows the application of adjustable electric fields to improve ion transmission and selection.
Furthermore, we are developing multi-laser configurations for LILBID-MS to better control the ionization mechanism and enhance sensitivity while preserving structural information on fragile biomolecular complexes.
Our group continuously develops new approaches to further improve existing methods and instrumentation for native mass spectrometry. For example, we are implementing Paul trap–based strategies to levitate microdroplets, enabling the investigation of fast, time-resolved reactions prior to ionization in LILBID-MS experiments.
In addition, we are advancing spectral quality in ESI-MS through MS-SIEVE, a setup that allows the application of adjustable electric fields to improve ion transmission and selection.
Furthermore, we are developing multi-laser configurations for LILBID-MS to better control the ionization mechanism and enhance sensitivity while preserving structural information on fragile biomolecular complexes.
